Data Portal for Single Cell Sequencing
Build an interactive viewer for single-cell RNA sequencing data using public datasets from the CZ CELLxGENE data portal.
Organise project
-
Create a folder for your project on your computer
-
Create a data folder outside of your project folder
Get single-cell data
-
Browse the CELLxGENE collections and find a dataset. For this example we use the Allen Institute Adult Human Brain Atlas - specifically the midbrain (substantia nigra) snRNA-seq dataset.
-
Download the
.h5adfile to your laptop (it is 416.2 MB). The dataset contains 59,505 cells x 58,232 genes with 11 cell types across 3 donors. Key metadata columns includecell_type,cluster_id,supercluster_term,donor_id, and UMAP coordinates inobsm['X_UMAP'].
Get prompting!
- In
Planmode, type:
Build an interactive viewer for my single-cell RNA data. The file is at [path/to/data.h5ad].
Result
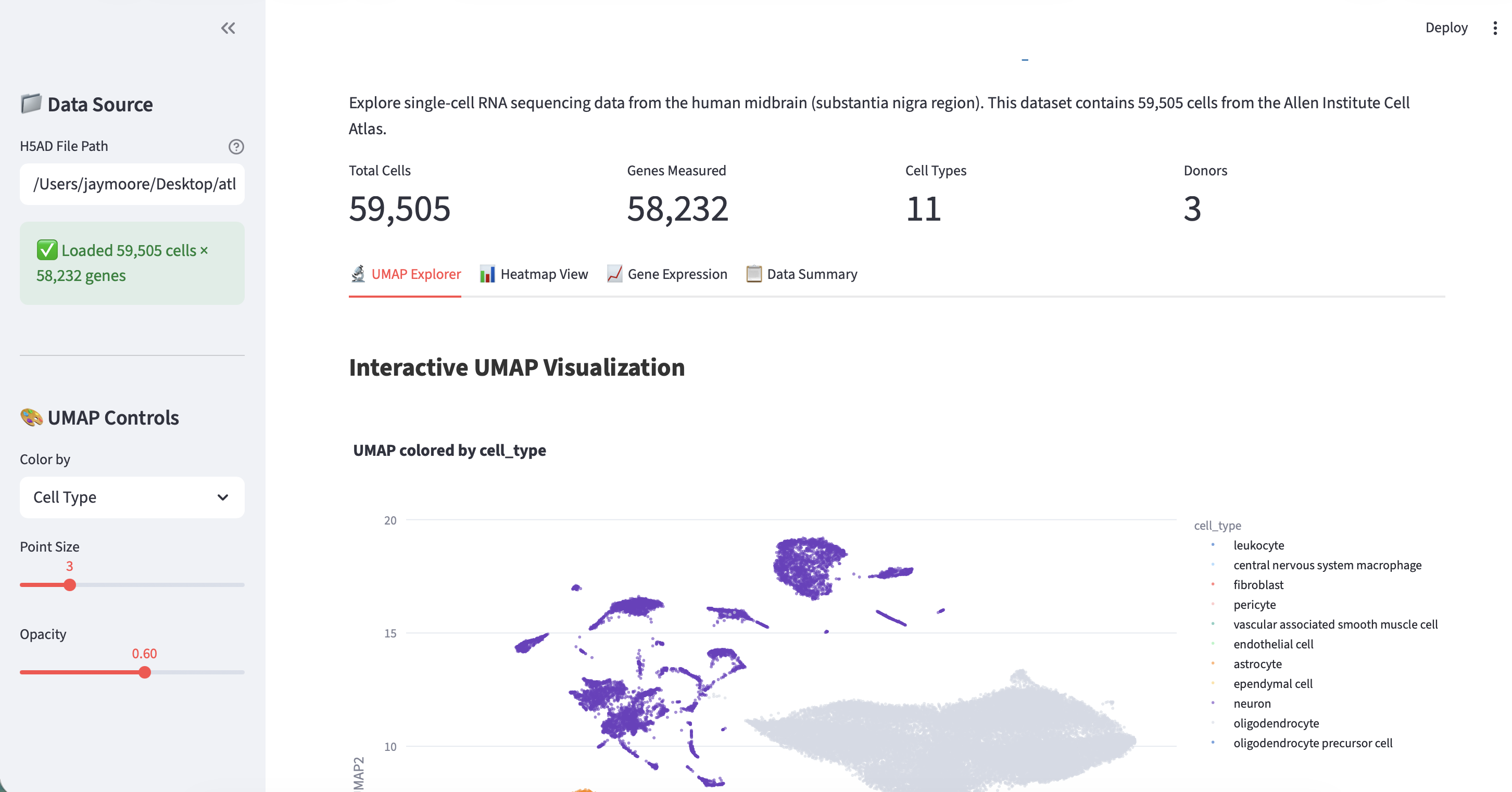
Publish to GitHub
Once your app is working, you can save it to GitHub:
Install GitHub CLI
- Download GitHub CLI
- Authenticate:
gh auth login
Upload your code
In chat mode, ask:
Push my project to a new GitHub repository called [your-repo-name]
Or create a repo first:
Create a new private GitHub repository called [your-repo-name], then push my code to it
Keep your context clean while working
Long sessions building and debugging your app will fill the context window quickly. Before pushing to GitHub, read the Managing Context tutorial to learn how to use /compact, maintain a PLAN.md, and avoid hallucinations on longer projects.