Infer species
Author: Brian M. Schilder
Author: Brian M. Schilder
Most recent update: Apr-29-2026
Source: Most recent update: Apr-29-2026
vignettes/infer_species.Rmd
infer_species.RmdInstallation
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
# orthogene is only available on Bioconductor>=3.14
if(BiocManager::version()<"3.14")
BiocManager::install(update = TRUE, ask = FALSE)
BiocManager::install("orthogene")Introduction
It’s not always clear whether a dataset is using the original species gene names, human gene names, or some other species’ gene names.
infer_species takes a list/matrix/data.frame with genes
and infers the species that they best match to!
For the sake of speed, the genes extracted from gene_df
are tested against genomes from only the following 6
test_species by default: - human - monkey - rat - mouse -
zebrafish - fly
However, you can supply your own list of test_species,
which will be automatically be mapped and standardised using
map_species.
Examples
Mouse genes
Infer the species
matches <- orthogene::infer_species(gene_df = exp_mouse,
method = method)## Preparing gene_df.## sparseMatrix format detected.## Extracting genes from rownames.## 15,259 genes extracted.## Testing for gene overlap with: human## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: human## Common name mapping found for human## 1 organism identified from search: 9606## Using cached file: /github/home/.cache/R/orthogene/all_genes-9606-homologene.csv.gz## Returning all 19,129 genes from human.## Testing for gene overlap with: monkey## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: monkey## Common name mapping found for monkey## 1 organism identified from search: 9544## Using cached file: /github/home/.cache/R/orthogene/all_genes-9544-homologene.csv.gz## Returning all 16,843 genes from monkey.## Testing for gene overlap with: rat## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: rat## Common name mapping found for rat## 1 organism identified from search: 10116## Using cached file: /github/home/.cache/R/orthogene/all_genes-10116-homologene.csv.gz## Returning all 20,616 genes from rat.## Testing for gene overlap with: mouse## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: mouse## Common name mapping found for mouse## 1 organism identified from search: 10090## Using cached file: /github/home/.cache/R/orthogene/all_genes-10090-homologene.csv.gz## Returning all 21,207 genes from mouse.## Testing for gene overlap with: zebrafish## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: zebrafish## Common name mapping found for zebrafish## 1 organism identified from search: 7955## Using cached file: /github/home/.cache/R/orthogene/all_genes-7955-homologene.csv.gz## Returning all 20,897 genes from zebrafish.## Testing for gene overlap with: fly## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: fly## Common name mapping found for fly## 1 organism identified from search: 7227## Using cached file: /github/home/.cache/R/orthogene/all_genes-7227-homologene.csv.gz## Returning all 8,438 genes from fly.## Top match:
## - species: mouse
## - percent_match: 92%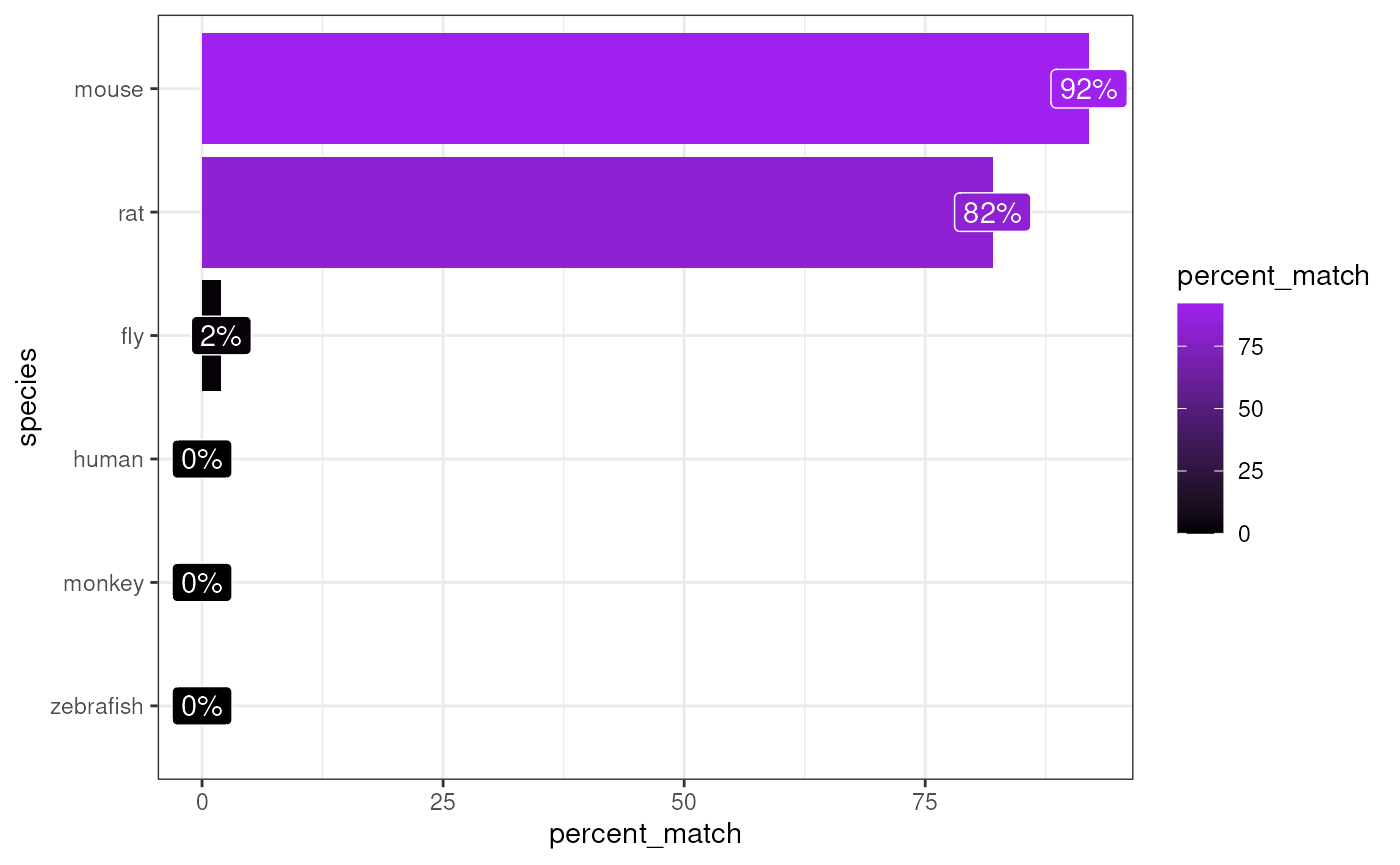
Rat genes
Create example data
To create an example dataset, turn the gene names into rat genes.
exp_rat <- orthogene::convert_orthologs(gene_df = exp_mouse,
input_species = "mouse",
output_species = "rat",
method = method)## Preparing gene_df.## sparseMatrix format detected.## Extracting genes from rownames.## 15,259 genes extracted.## Converting mouse ==> rat orthologs using: homologene## Retrieving all organisms available in homologene.## Mapping species name: mouse## Common name mapping found for mouse## 1 organism identified from search: 10090## Retrieving all organisms available in homologene.## Mapping species name: rat## Common name mapping found for rat## 1 organism identified from search: 10116## Checking for genes without orthologs in rat.## Extracting genes from input_gene.## 13,812 genes extracted.## Extracting genes from ortholog_gene.## 13,812 genes extracted.## Checking for genes without 1:1 orthologs.## Dropping 486 genes that have multiple input_gene per ortholog_gene (many:1).## Dropping 148 genes that have multiple ortholog_gene per input_gene (1:many).## Filtering gene_df with gene_map## Setting ortholog_gene to rownames.##
## =========== REPORT SUMMARY ===========## Total genes dropped after convert_orthologs :
## 2,322 / 15,259 (15%)## Total genes remaining after convert_orthologs :
## 12,937 / 15,259 (85%)Infer the species
matches <- orthogene::infer_species(gene_df = exp_rat,
method = method)## Preparing gene_df.## sparseMatrix format detected.## Extracting genes from rownames.## 12,937 genes extracted.## Testing for gene overlap with: human## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: human## Common name mapping found for human## 1 organism identified from search: 9606## Using cached file: /github/home/.cache/R/orthogene/all_genes-9606-homologene.csv.gz## Returning all 19,129 genes from human.## Testing for gene overlap with: monkey## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: monkey## Common name mapping found for monkey## 1 organism identified from search: 9544## Using cached file: /github/home/.cache/R/orthogene/all_genes-9544-homologene.csv.gz## Returning all 16,843 genes from monkey.## Testing for gene overlap with: rat## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: rat## Common name mapping found for rat## 1 organism identified from search: 10116## Using cached file: /github/home/.cache/R/orthogene/all_genes-10116-homologene.csv.gz## Returning all 20,616 genes from rat.## Testing for gene overlap with: mouse## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: mouse## Common name mapping found for mouse## 1 organism identified from search: 10090## Using cached file: /github/home/.cache/R/orthogene/all_genes-10090-homologene.csv.gz## Returning all 21,207 genes from mouse.## Testing for gene overlap with: zebrafish## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: zebrafish## Common name mapping found for zebrafish## 1 organism identified from search: 7955## Using cached file: /github/home/.cache/R/orthogene/all_genes-7955-homologene.csv.gz## Returning all 20,897 genes from zebrafish.## Testing for gene overlap with: fly## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: fly## Common name mapping found for fly## 1 organism identified from search: 7227## Using cached file: /github/home/.cache/R/orthogene/all_genes-7227-homologene.csv.gz## Returning all 8,438 genes from fly.## Top match:
## - species: rat
## - percent_match: 100%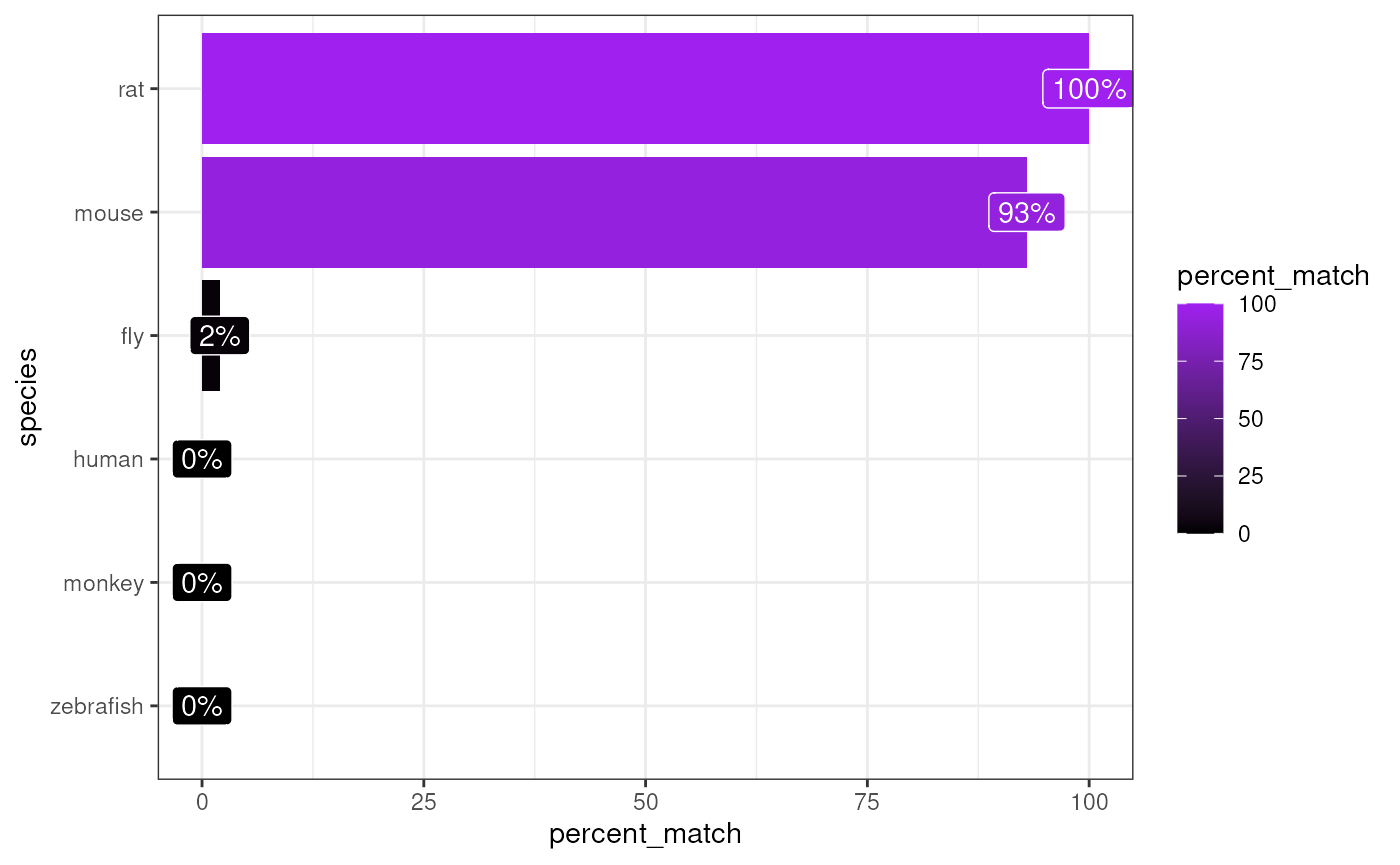
Human genes
Create example data
To create an example dataset, turn the gene names into human genes.
exp_human <- orthogene::convert_orthologs(gene_df = exp_mouse,
input_species = "mouse",
output_species = "human",
method = method)## Preparing gene_df.## sparseMatrix format detected.## Extracting genes from rownames.## 15,259 genes extracted.## Converting mouse ==> human orthologs using: homologene## Retrieving all organisms available in homologene.## Mapping species name: mouse## Common name mapping found for mouse## 1 organism identified from search: 10090## Retrieving all organisms available in homologene.## Mapping species name: human## Common name mapping found for human## 1 organism identified from search: 9606## Checking for genes without orthologs in human.## Extracting genes from input_gene.## 13,416 genes extracted.## Extracting genes from ortholog_gene.## 13,416 genes extracted.## Checking for genes without 1:1 orthologs.## Dropping 46 genes that have multiple input_gene per ortholog_gene (many:1).## Dropping 56 genes that have multiple ortholog_gene per input_gene (1:many).## Filtering gene_df with gene_map## Setting ortholog_gene to rownames.##
## =========== REPORT SUMMARY ===========## Total genes dropped after convert_orthologs :
## 2,016 / 15,259 (13%)## Total genes remaining after convert_orthologs :
## 13,243 / 15,259 (87%)Infer the species
matches <- orthogene::infer_species(gene_df = exp_human,
method = method)## Preparing gene_df.## sparseMatrix format detected.## Extracting genes from rownames.## 13,243 genes extracted.## Testing for gene overlap with: human## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: human## Common name mapping found for human## 1 organism identified from search: 9606## Using cached file: /github/home/.cache/R/orthogene/all_genes-9606-homologene.csv.gz## Returning all 19,129 genes from human.## Testing for gene overlap with: monkey## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: monkey## Common name mapping found for monkey## 1 organism identified from search: 9544## Using cached file: /github/home/.cache/R/orthogene/all_genes-9544-homologene.csv.gz## Returning all 16,843 genes from monkey.## Testing for gene overlap with: rat## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: rat## Common name mapping found for rat## 1 organism identified from search: 10116## Using cached file: /github/home/.cache/R/orthogene/all_genes-10116-homologene.csv.gz## Returning all 20,616 genes from rat.## Testing for gene overlap with: mouse## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: mouse## Common name mapping found for mouse## 1 organism identified from search: 10090## Using cached file: /github/home/.cache/R/orthogene/all_genes-10090-homologene.csv.gz## Returning all 21,207 genes from mouse.## Testing for gene overlap with: zebrafish## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: zebrafish## Common name mapping found for zebrafish## 1 organism identified from search: 7955## Using cached file: /github/home/.cache/R/orthogene/all_genes-7955-homologene.csv.gz## Returning all 20,897 genes from zebrafish.## Testing for gene overlap with: fly## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: fly## Common name mapping found for fly## 1 organism identified from search: 7227## Using cached file: /github/home/.cache/R/orthogene/all_genes-7227-homologene.csv.gz## Returning all 8,438 genes from fly.## Top match:
## - species: human
## - percent_match: 100%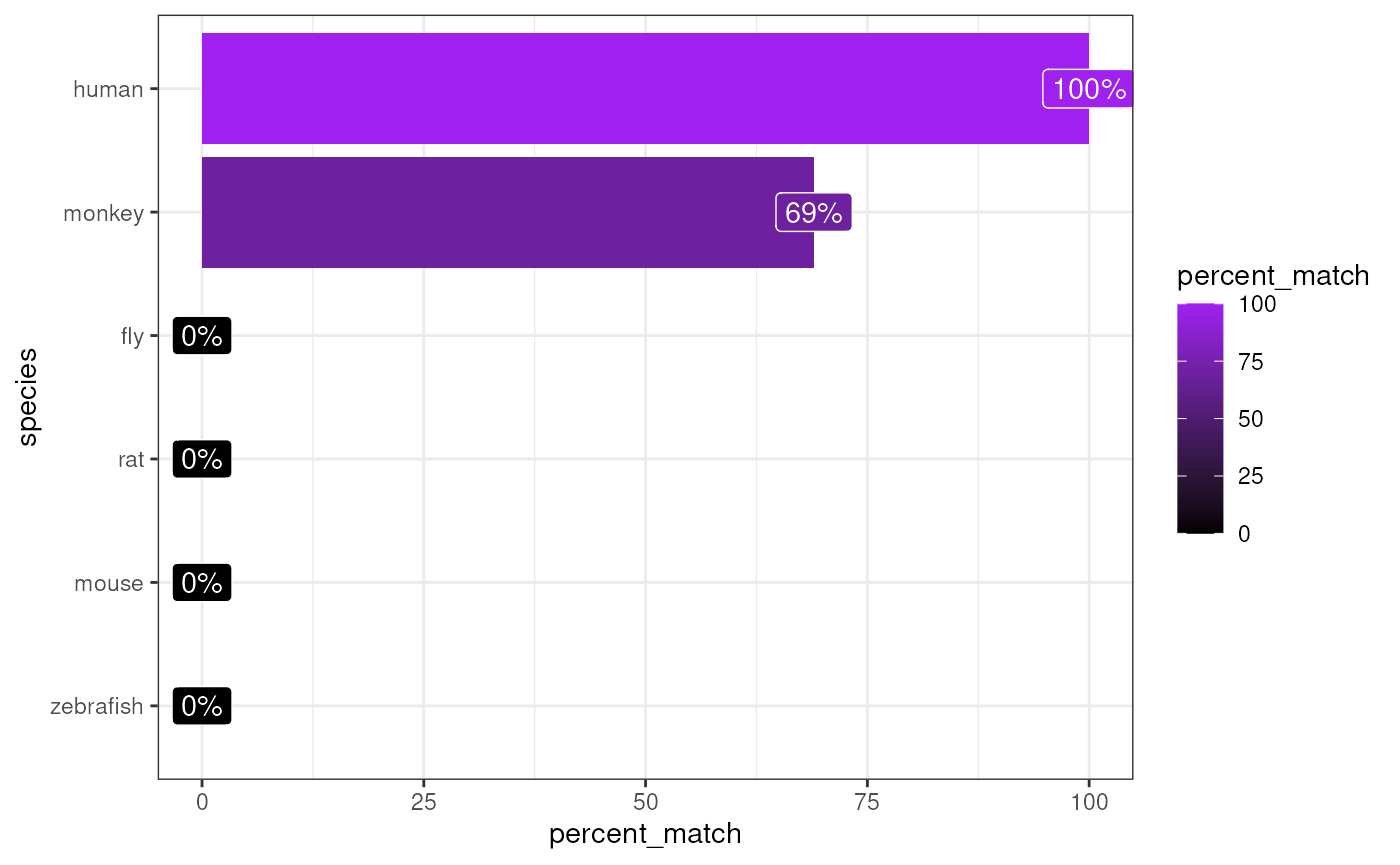
Additional test_species
You can even supply test_species with the name of one of
the R packages that orthogene gets orthologs from. This
will test against all species available in that particular R
package.
For example, by setting test_species="homologene" we
automatically test for % gene matches in each of the 20+ species
available in homologene.
matches <- orthogene::infer_species(gene_df = exp_human,
test_species = method,
method = method)## Retrieving all organisms available in homologene.## Preparing gene_df.## sparseMatrix format detected.## Extracting genes from rownames.## 13,243 genes extracted.## Testing for gene overlap with: Mus musculus## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Mus musculus## 1 organism identified from search: 10090## Using cached file: /github/home/.cache/R/orthogene/all_genes-10090-homologene.csv.gz## Returning all 21,207 genes from Mus musculus.## Testing for gene overlap with: Rattus norvegicus## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Rattus norvegicus## 1 organism identified from search: 10116## Using cached file: /github/home/.cache/R/orthogene/all_genes-10116-homologene.csv.gz## Returning all 20,616 genes from Rattus norvegicus.## Testing for gene overlap with: Kluyveromyces lactis## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Kluyveromyces lactis## 1 organism identified from search: 28985## Using cached file: /github/home/.cache/R/orthogene/all_genes-28985-homologene.csv.gz## Returning all 4,283 genes from Kluyveromyces lactis.## Testing for gene overlap with: Magnaporthe oryzae## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Magnaporthe oryzae## 1 organism identified from search: 318829## Using cached file: /github/home/.cache/R/orthogene/all_genes-318829-homologene.csv.gz## Returning all 6,598 genes from Magnaporthe oryzae.## Testing for gene overlap with: Eremothecium gossypii## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Eremothecium gossypii## 1 organism identified from search: 33169## Using cached file: /github/home/.cache/R/orthogene/all_genes-33169-homologene.csv.gz## Returning all 3,874 genes from Eremothecium gossypii.## Testing for gene overlap with: Arabidopsis thaliana## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Arabidopsis thaliana## 1 organism identified from search: 3702## Using cached file: /github/home/.cache/R/orthogene/all_genes-3702-homologene.csv.gz## Returning all 19,143 genes from Arabidopsis thaliana.## Testing for gene overlap with: Oryza sativa## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Oryza sativa## 1 organism identified from search: 4530## Using cached file: /github/home/.cache/R/orthogene/all_genes-4530-homologene.csv.gz## Returning all 16,112 genes from Oryza sativa.## Testing for gene overlap with: Schizosaccharomyces pombe## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Schizosaccharomyces pombe## 1 organism identified from search: 4896## Using cached file: /github/home/.cache/R/orthogene/all_genes-4896-homologene.csv.gz## Returning all 3,018 genes from Schizosaccharomyces pombe.## Testing for gene overlap with: Saccharomyces cerevisiae## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Saccharomyces cerevisiae## 1 organism identified from search: 4932## Using cached file: /github/home/.cache/R/orthogene/all_genes-4932-homologene.csv.gz## Returning all 4,579 genes from Saccharomyces cerevisiae.## Testing for gene overlap with: Neurospora crassa## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Neurospora crassa## 1 organism identified from search: 5141## Using cached file: /github/home/.cache/R/orthogene/all_genes-5141-homologene.csv.gz## Returning all 5,807 genes from Neurospora crassa.## Testing for gene overlap with: Caenorhabditis elegans## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Caenorhabditis elegans## 1 organism identified from search: 6239## Using cached file: /github/home/.cache/R/orthogene/all_genes-6239-homologene.csv.gz## Returning all 7,575 genes from Caenorhabditis elegans.## Testing for gene overlap with: Anopheles gambiae## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Anopheles gambiae## 1 organism identified from search: 7165## Using cached file: /github/home/.cache/R/orthogene/all_genes-7165-homologene.csv.gz## Returning all 8,428 genes from Anopheles gambiae.## Testing for gene overlap with: Drosophila melanogaster## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Drosophila melanogaster## 1 organism identified from search: 7227## Using cached file: /github/home/.cache/R/orthogene/all_genes-7227-homologene.csv.gz## Returning all 8,438 genes from Drosophila melanogaster.## Testing for gene overlap with: Danio rerio## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Danio rerio## 1 organism identified from search: 7955## Using cached file: /github/home/.cache/R/orthogene/all_genes-7955-homologene.csv.gz## Returning all 20,897 genes from Danio rerio.## Testing for gene overlap with: Xenopus (Silurana) tropicalis## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Xenopus (Silurana) tropicalis## 1 organism identified from search: 8364## Using cached file: /github/home/.cache/R/orthogene/all_genes-8364-homologene.csv.gz## Returning all 18,446 genes from Xenopus (Silurana) tropicalis.## Testing for gene overlap with: Gallus gallus## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Gallus gallus## 1 organism identified from search: 9031## Using cached file: /github/home/.cache/R/orthogene/all_genes-9031-homologene.csv.gz## Returning all 14,600 genes from Gallus gallus.## Testing for gene overlap with: Macaca mulatta## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Macaca mulatta## 1 organism identified from search: 9544## Using cached file: /github/home/.cache/R/orthogene/all_genes-9544-homologene.csv.gz## Returning all 16,843 genes from Macaca mulatta.## Testing for gene overlap with: Pan troglodytes## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Pan troglodytes## 1 organism identified from search: 9598## Using cached file: /github/home/.cache/R/orthogene/all_genes-9598-homologene.csv.gz## Returning all 18,730 genes from Pan troglodytes.## Testing for gene overlap with: Homo sapiens## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Homo sapiens## 1 organism identified from search: 9606## Using cached file: /github/home/.cache/R/orthogene/all_genes-9606-homologene.csv.gz## Returning all 19,129 genes from Homo sapiens.## Testing for gene overlap with: Canis lupus familiaris## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Canis lupus familiaris## 1 organism identified from search: 9615## Using cached file: /github/home/.cache/R/orthogene/all_genes-9615-homologene.csv.gz## Returning all 18,117 genes from Canis lupus familiaris.## Testing for gene overlap with: Bos taurus## Retrieving all genes using: homologene.## Retrieving all organisms available in homologene.## Mapping species name: Bos taurus## 1 organism identified from search: 9913## Using cached file: /github/home/.cache/R/orthogene/all_genes-9913-homologene.csv.gz## Returning all 18,797 genes from Bos taurus.## Top match:
## - species: Homo sapiens
## - percent_match: 100%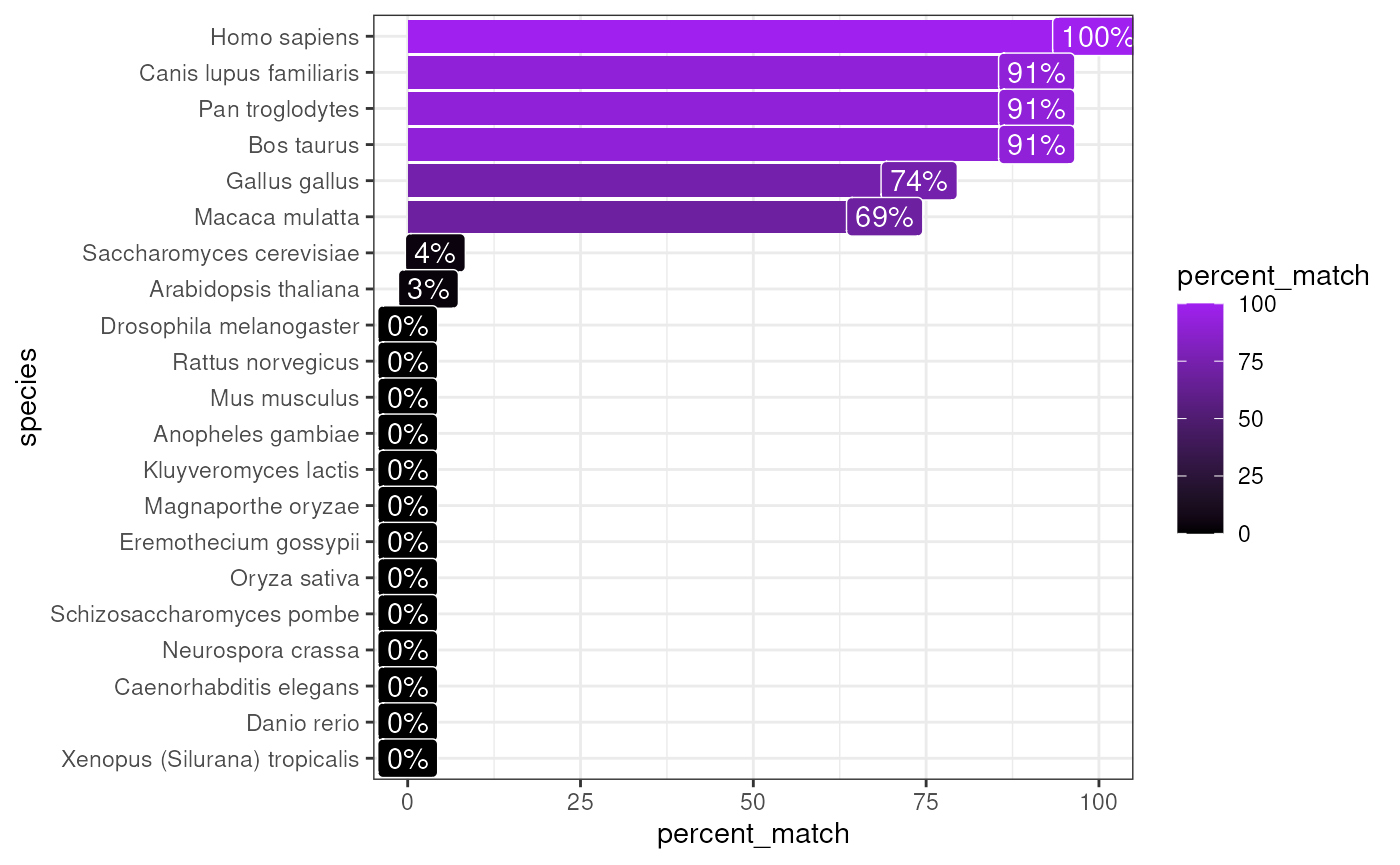
Session Info
utils::sessionInfo()## R version 4.6.0 (2026-04-24)
## Platform: x86_64-pc-linux-gnu
## Running under: Ubuntu 24.04.4 LTS
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
## LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
##
## locale:
## [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
## [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
## [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
##
## time zone: UTC
## tzcode source: system (glibc)
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] orthogene_1.19.3 BiocStyle_2.40.0
##
## loaded via a namespace (and not attached):
## [1] ggiraph_0.9.6 tidyselect_1.2.1
## [3] viridisLite_0.4.3 dplyr_1.2.1
## [5] farver_2.1.2 R.utils_2.13.0
## [7] S7_0.2.2 fastmap_1.2.0
## [9] lazyeval_0.2.3 homologene_1.4.68.19.3.27
## [11] fontquiver_0.2.1 digest_0.6.39
## [13] lifecycle_1.0.5 tidytree_0.4.7
## [15] magrittr_2.0.5 compiler_4.6.0
## [17] rlang_1.2.0 sass_0.4.10
## [19] tools_4.6.0 yaml_2.3.12
## [21] data.table_1.18.2.1 knitr_1.51
## [23] ggsignif_0.6.4 labeling_0.4.3
## [25] htmlwidgets_1.6.4 RColorBrewer_1.1-3
## [27] aplot_0.2.9 abind_1.4-8
## [29] babelgene_22.9 withr_3.0.2
## [31] purrr_1.2.2 R.oo_1.27.1
## [33] desc_1.4.3 grid_4.6.0
## [35] ggpubr_0.6.3 gdtools_0.5.0
## [37] ggplot2_4.0.3 scales_1.4.0
## [39] MASS_7.3-65 cli_3.6.6
## [41] rmarkdown_2.31 ragg_1.5.2
## [43] treeio_1.36.0 generics_0.1.4
## [45] otel_0.2.0 ggtree_4.2.0
## [47] httr_1.4.8 gprofiler2_0.2.4
## [49] ape_5.8-1 cachem_1.1.0
## [51] parallel_4.6.0 ggplotify_0.1.3
## [53] BiocManager_1.30.27 yulab.utils_0.2.4
## [55] vctrs_0.7.3 Matrix_1.7-5
## [57] jsonlite_2.0.0 fontBitstreamVera_0.1.1
## [59] carData_3.0-6 bookdown_0.46
## [61] car_3.1-5 gridGraphics_0.5-1
## [63] patchwork_1.3.2 rstatix_0.7.3
## [65] Formula_1.2-5 systemfonts_1.3.2
## [67] plotly_4.12.0 tidyr_1.3.2
## [69] jquerylib_0.1.4 glue_1.8.1
## [71] pkgdown_2.2.0 gtable_0.3.6
## [73] tibble_3.3.1 pillar_1.11.1
## [75] rappdirs_0.3.4 htmltools_0.5.9
## [77] R6_2.6.1 textshaping_1.0.5
## [79] evaluate_1.0.5 lattice_0.22-9
## [81] R.methodsS3_1.8.2 backports_1.5.1
## [83] broom_1.0.12 ggfun_0.2.0
## [85] fontLiberation_0.1.0 bslib_0.10.0
## [87] Rcpp_1.1.1-1.1 nlme_3.1-169
## [89] xfun_0.57 fs_2.1.0
## [91] pkgconfig_2.0.3