Harmonizing Cell-type Labels
Author: Brian M. Schilder
Author: Brian M. Schilder
Most recent update: Jan-26-2026
Source: Most recent update: Jan-26-2026
vignettes/harmonise_celltypes.Rmd
harmonise_celltypes.RmdIntroduction
Cell-type labels from different datasets are often inconsistent. Authors use different naming conventions, levels of granularity, and terminology. For example:
- “Purkinje neurons” vs “Purkinje cells” vs “PCs”
- “Astrocytes” vs “Astrocyte-like cells” vs “GFAP+ glia”
- “Inhibitory neurons” vs “GABAergic neurons” vs “Interneurons”
scNLP provides NLP-based approaches to identify
semantically similar labels and harmonize cell-type annotations across
datasets without requiring manual curation.
Example Data
We’ll use the included pseudo-bulk dataset containing samples from multiple studies with heterogeneous cell-type labels:
data("pseudo_seurat")
# View the diversity of labels and sources
label_summary <- table(
Dataset = pseudo_seurat$batch,
Species = pseudo_seurat$species
)
print(label_summary)## Species
## Dataset fly human mouse zebrafish
## AIBS 0 6 0 0
## Aerts2020 116 0 0 0
## BlueLake2018_FrontalCortexOnly 0 24 0 0
## BlueLake2018_VisualCortexOnly 0 28 0 0
## DRONC_human 0 15 0 0
## DRONC_mouse 0 0 22 0
## LaManno2020 0 0 206 0
## Raj2020 0 0 0 222
## Zeisel2016 0 0 48 0
## Zeisel2018 0 0 37 0
## descartes_SampledData 0 77 0 0## [1] 608## [1] ASC1 ASC2 END exCA1 exCA3 exDG exPFC1 exPFC2 GABA1 GABA2
## [11] MG NSC ODC1 ODC2 OPC ChP exPFC3 exPFC4 exPFC5 exPFC6
## 608 Levels: A AFP_ALB_positive_cells ASC1 ASC2 ... ventral_midbrainStrategy 1: TF-IDF Clustering
TF-IDF analysis identifies enriched terms within each cluster. Cells with similar enriched terms likely represent the same cell type, regardless of the exact label used.
Compute TF-IDF
# Run TF-IDF on the existing clusters
result <- run_tfidf(
obj = pseudo_seurat,
reduction = "umap",
cluster_var = "cluster",
label_var = "celltype"
)## Extracting obsm from Seurat: umap## + Dropping 2 conflicting obs variables: UMAP.1, UMAP.2## Loading required namespace: tidytext## Setting cell metadata (obs) in obj.Visualize Enriched Terms
The scatter plot shows cells colored by cluster, with enriched terms labeled at cluster centers:
res <- plot_tfidf(
obj = pseudo_seurat,
label_var = "celltype",
cluster_var = "cluster",
show_plot = TRUE
)## Extracting obsm from Seurat: umap## + Dropping 2 conflicting obs variables: UMAP.1, UMAP.2## Setting cell metadata (obs) in obj.## Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
## ℹ Please use tidy evaluation idioms with `aes()`.
## ℹ See also `vignette("ggplot2-in-packages")` for more information.
## ℹ The deprecated feature was likely used in the scNLP package.
## Please report the issue at <https://github.com/neurogenomics/scNLP/issues>.
## This warning is displayed once per session.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.## Warning in ggplot2::geom_point(ggplot2::aes_string(color = color_var, size =
## size_var, : Ignoring unknown aesthetics: label## Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
## ℹ Please use `linewidth` instead.
## ℹ The deprecated feature was likely used in the scNLP package.
## Please report the issue at <https://github.com/neurogenomics/scNLP/issues>.
## This warning is displayed once per session.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.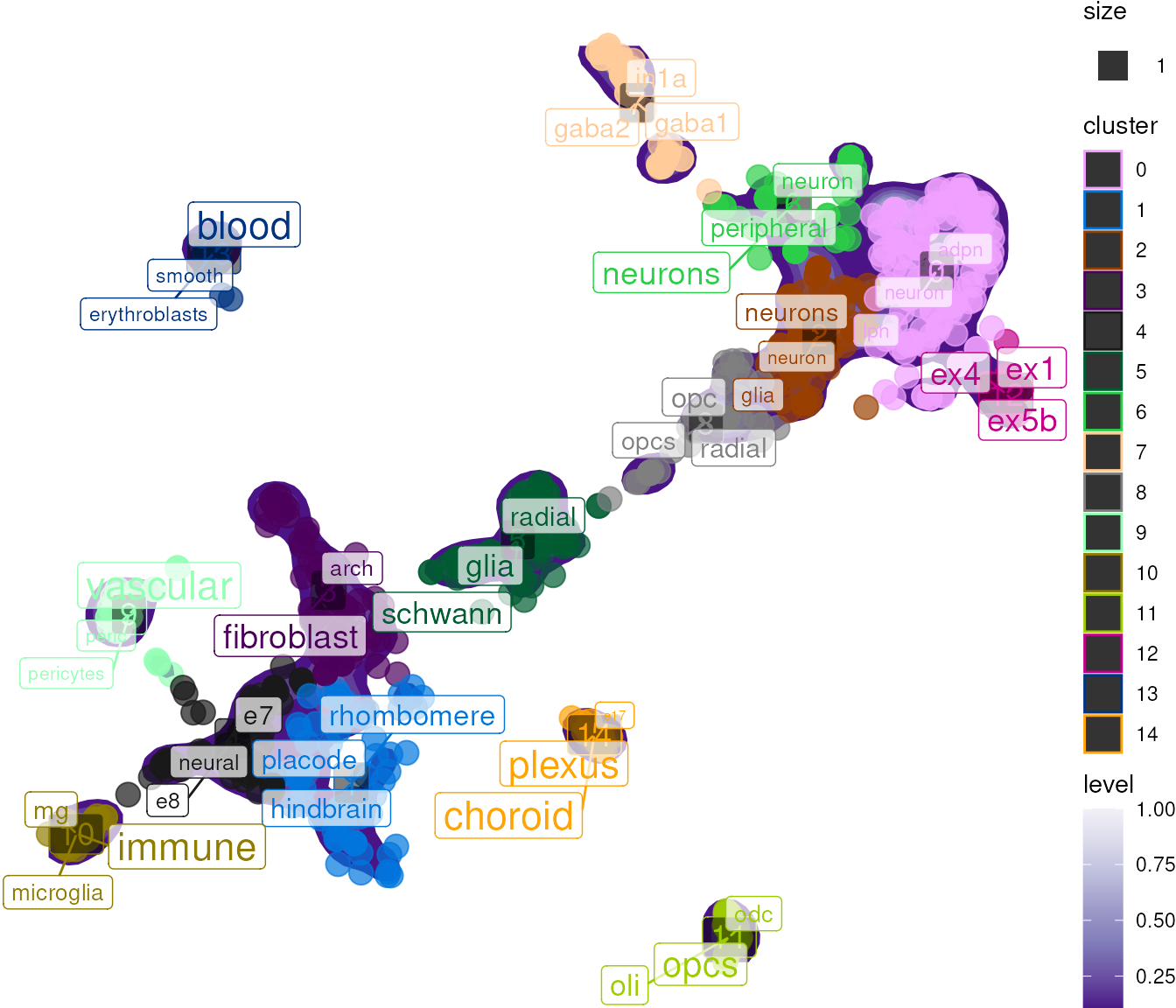
Examine TF-IDF Results
The TF-IDF results reveal which terms are most distinctive for each cluster:
# Get the full TF-IDF results
tfidf_df <- res$tfidf_df
# Top terms per cluster
top_terms <- tfidf_df |>
dplyr::group_by(cluster) |>
dplyr::slice_max(order_by = tf_idf, n = 3) |>
dplyr::select(cluster, word, tf_idf)
print(top_terms, n = 30)## # A tibble: 44 × 3
## # Groups: cluster [15]
## cluster word tf_idf
## <fct> <chr> <dbl>
## 1 0 lpn 0.0528
## 2 0 adpn 0.0523
## 3 0 neuron 0.0428
## 4 1 rhombomere 0.154
## 5 1 placode 0.116
## 6 1 hindbrain 0.114
## 7 2 neurons 0.101
## 8 2 glia 0.0587
## 9 2 neuron 0.0419
## 10 3 fibroblast 0.249
## 11 3 arch 0.0653
## 12 4 e7 0.112
## 13 4 e8 0.0612
## 14 4 neural 0.0596
## 15 5 glia 0.198
## 16 5 schwann 0.182
## 17 5 radial 0.112
## 18 6 neurons 0.239
## 19 6 peripheral 0.104
## 20 6 neuron 0.0704
## 21 7 gaba1 0.120
## 22 7 gaba2 0.120
## 23 7 in1a 0.120
## 24 8 opc 0.143
## 25 8 radial 0.119
## 26 8 opcs 0.102
## 27 9 vascular 0.528
## 28 9 peric 0.0423
## 29 9 pericytes 0.0423
## 30 10 immune 0.521
## # ℹ 14 more rowsClusters sharing similar enriched terms (e.g., “neuron”, “astrocyte”, “oligodendrocyte”) likely contain the same cell types from different datasets.
Strategy 2: Neighbor-based Harmonization
The search_neighbors() function finds cells with similar
expression profiles, allowing you to identify which labels from
different datasets correspond to the same cell types.
Find Similar Cells
Search for cells similar to those labeled “purkinje”:
purkinje_neighbors <- search_neighbors(
seurat = pseudo_seurat,
var1_search = "purkinje",
max_neighbors = 10
)## No variable features detected. Computing## No PCA detected. Computing## Centering and scaling data matrix## PC_ 1
## Positive: FTMT, ACTG1, ALAS2, HSPA1L, NME1-NME2, POTEI, OTOP2, RAPSN, BEST2, DPEP2
## HMGB2, NAPRT1, KRT8, PPIB, DSCC1, POU4F3, CCDC102A, GAPDHS, CHST6, AGXT2
## LYZL2, MTMR8, ACTG2, ACPL2, BANF1, PPAPDC2, HTR1A, IFI30, CYBRD1, LHX8
## Negative: FAIM2, CAMK2A, SCN1A, CAMK2B, FRRS1L, UNC80, PHYHIP, RASGRF2, CCK, GRIA2
## STXBP5L, ARPP21, SLC12A5, DIRAS2, RYR2, SLC4A10, KCNT1, GRM5, CAMKV, KIAA1211L
## GABRA4, GABRA1, SV2B, CX3CL1, AK5, PNMA2, JPH4, DGKG, GPR158, KCNC2
## PC_ 2
## Positive: CAMKK1, DGKQ, NT5DC3, CA7, ABCG4, HTR1A, C5orf28, OTOP2, HYKK, DPEP2
## CHST6, POTEI, SLC8A3, SLC38A11, ADRA2A, MPPED1, MTMR8, HTR7, CACNA1B, PPAPDC2
## C2orf69, GRIK1, IFI30, STK32B, RASL10B, SLC24A4, FAXDC2, ADCY3, ACSS2, ANKRD29
## Negative: RAN, HSP90AA1, H2AFZ, HNRNPAB, CCT5, NPM1, GNG5, DBI, HMGB2, ITM2B
## ATP6V1G1, SERPINH1, CIRBP, CD63, NDUFA6, MDK, JUN, MYL12B, SPARC, NPC2
## GLUL, ID3, EEF1A1, VIM, CLIC1, COX6B1, LDHA, DDAH2, ENO1, CNN3
## PC_ 3
## Positive: ADGRL2, AC011288.2, RP11-420N3.3, RP11-191L9.4, NRXN3, PLPPR1, RP11-123O10.4, ZNF385D, AC114765.1, NWD2
## RBFOX3, MIR137HG, MIR325HG, SGOL1-AS1, POU6F2, ANKRD18A, LY86-AS1, LINC01197, DGCR5, DPY19L1P1
## MIR4300HG, AQP4-AS1, HPSE2, LINC00632, NLGN4X, AC067956.1, PWRN1, LINC00599, CABP1, LINC01158
## Negative: KRTCAP2, APOE, C20orf24, PDIA6, PGLS, GNG11, S100A13, HIST1H2BI, ISCA2, GSTM5
## LAPTM4A, CST3, TMEM176B, KLF4, PDLIM2, CAP1, S100A16, APRT, CYR61, FAIM
## IFITM3, CDKN1A, KLF2, CLIC1, ARPC1B, IER2, S100A1, CMTM5, FXYD1, TCN2
## PC_ 4
## Positive: RESP18, CTXN2, ATP6V1G2, GNG13, DISP2, C15orf59, CCDC85A, GNG3, SYNGR3, RGS8
## VWA5B2, C1QL3, HPCA, TUBB3, CALB1, SNCB, HTR3A, ARHGDIG, L1CAM, NAP1L5
## PCDH20, HMP19, DBNDD2, NPAS4, FABP3, CALY, FAM43B, CKMT1B, LOC728392, LTK
## Negative: PTPN18, SLCO1A2, LINC00639, INPP5D, IFI44, LYN, DISC1, NEAT1, NRGN, CMYA5
## IFI44L, GALNT15, PARP14, AC012593.1, AQP4-AS1, MSR1, MT2A, ISG15, SHROOM4, CABP1
## UACA, KCNQ1OT1, PART1, CNDP1, FAM153B, DGCR5, SOX2-OT, LINC00844, ADGRG1, LINC00599
## PC_ 5
## Positive: MEST, IGFBP2, CNN3, FBXL7, NNAT, TUBB2B, GPC3, VIM, NKAIN4, ID1
## BMP7, CSRP2, NDN, DDAH2, GPX8, IGFBPL1, MARCKSL1, GSTM3, FBLN1, PARD3
## MFAP4, PTN, FABP7, COPS6, CTNNA2, ZBTB20, BEX1, CD81, ENO1, NPAS3
## Negative: C1QB, FCGR2A, MS4A6A, TYROBP, C1QC, AIF1, C1QA, CSF1R, CD86, MRC1
## MS4A7, CTSS, CCL24, FCER1G, CD53, CD14, FCGR1A, PLEK, C3AR1, LYZ
## FCGR2B, CX3CR1, CCL3L3, CCL2, CCR1, CD68, C5AR1, PF4, HPGDS, LY86## No graphs detected. Computing.## Computing nearest neighbor graph## Computing SNN## Using graph: RNA_snn## + Filtering results by `var1_search`: purkinje## + 3 entries matching `var1_search` identified.## + Adding original names to results## + Returning 32 pair-wise similarities.
head(purkinje_neighbors)## Var1
## <char>
## 1: human.descartes_SampledData.Purkinje_neurons
## 2: human.descartes_SampledData.Purkinje_neurons
## 3: human.descartes_SampledData.Purkinje_neurons
## 4: human.descartes_SampledData.Purkinje_neurons
## 5: zebrafish.Raj2020.purkinje_neurons
## 6: zebrafish.Raj2020.purkinje_neurons
## Var2 similarity
## <char> <num>
## 1: human.descartes_SampledData.SATB2_LRRC7_positive_cells 0.9047619
## 2: human.descartes_SampledData.Oligodendrocytes 0.8181818
## 3: human.descartes_SampledData.Granule_neurons 0.8181818
## 4: human.descartes_SampledData.ENS_neurons 0.8181818
## 5: zebrafish.Raj2020.neurons._gabaergic._glutamatergic 0.8181818
## 6: zebrafish.Raj2020.purkinje_neurons_.gabaergic_... 0.8181818
## Var1_id
## <char>
## 1: human.descartes_SampledData.Purkinje_neurons
## 2: human.descartes_SampledData.Purkinje_neurons
## 3: human.descartes_SampledData.Purkinje_neurons
## 4: human.descartes_SampledData.Purkinje_neurons
## 5: zebrafish.Raj2020.purkinje_neurons
## 6: zebrafish.Raj2020.purkinje_neurons
## Var2_id
## <char>
## 1: human.descartes_SampledData.SATB2_LRRC7_positive_cells
## 2: human.descartes_SampledData.Oligodendrocytes
## 3: human.descartes_SampledData.Granule_neurons
## 4: human.descartes_SampledData.ENS_neurons
## 5: zebrafish.Raj2020.neurons._gabaergic._glutamatergic
## 6: zebrafish.Raj2020.purkinje_neurons_.gabaergic_...Cross-dataset Mapping
Examine which labels from other datasets map to Purkinje cells:
# Get unique neighbor labels
neighbor_labels <- unique(purkinje_neighbors$Var2)
cat("Labels similar to 'purkinje':\n")## Labels similar to 'purkinje':
print(neighbor_labels)## [1] "human.descartes_SampledData.SATB2_LRRC7_positive_cells"
## [2] "human.descartes_SampledData.Oligodendrocytes"
## [3] "human.descartes_SampledData.Granule_neurons"
## [4] "human.descartes_SampledData.ENS_neurons"
## [5] "zebrafish.Raj2020.neurons._gabaergic._glutamatergic"
## [6] "zebrafish.Raj2020.purkinje_neurons_.gabaergic_..."
## [7] "zebrafish.Raj2020.progenitors.differentiating_granule_cells_.hindbrain."
## [8] "zebrafish.Raj2020.ventral_habenula"
## [9] "zebrafish.Raj2020.purkinje_neurons"
## [10] "human.descartes_SampledData.Inhibitory_interneurons"
## [11] "human.descartes_SampledData.Excitatory_neurons"
## [12] "zebrafish.Raj2020.telencephalon_.pallium._neuron._glutamatergic"
## [13] "zebrafish.Raj2020.cranial_ganglion"
## [14] "zebrafish.Raj2020.neurons_.glutamatergic._midbrain."
## [15] "zebrafish.Raj2020.diencephalon.mid"
## [16] "zebrafish.Raj2020.neurons_.gabaergic_..._midbrain._optic_tectum."
## [17] "human.descartes_SampledData.SKOR2_NPSR1_positive_cells"
## [18] "zebrafish.Raj2020.retina_neurons_.horizontal.."
## [19] "zebrafish.Raj2020.neurons_.no_neurod1.neurod4._glutamatergic"
## [20] "human.descartes_SampledData.SLC24A4_PEX5L_positive_cells"
## [21] "human.descartes_SampledData.Inhibitory_neurons"
## [22] "human.descartes_SampledData.Amacrine_cells"
## [23] "human.descartes_SampledData.PDE11A_FAM19A2_positive_cells"Search Multiple Cell Types
You can search for multiple cell types to build a harmonization map:
# Search for astrocyte-like cells
astro_neighbors <- search_neighbors(
seurat = pseudo_seurat,
var1_search = "astro",
max_neighbors = 5
)## No variable features detected. Computing## No PCA detected. Computing## Centering and scaling data matrix## PC_ 1
## Positive: FTMT, ACTG1, ALAS2, HSPA1L, NME1-NME2, POTEI, OTOP2, RAPSN, BEST2, DPEP2
## HMGB2, NAPRT1, KRT8, PPIB, DSCC1, POU4F3, CCDC102A, GAPDHS, CHST6, AGXT2
## LYZL2, MTMR8, ACTG2, ACPL2, BANF1, PPAPDC2, HTR1A, IFI30, CYBRD1, LHX8
## Negative: FAIM2, CAMK2A, SCN1A, CAMK2B, FRRS1L, UNC80, PHYHIP, RASGRF2, CCK, GRIA2
## STXBP5L, ARPP21, SLC12A5, DIRAS2, RYR2, SLC4A10, KCNT1, GRM5, CAMKV, KIAA1211L
## GABRA4, GABRA1, SV2B, CX3CL1, AK5, PNMA2, JPH4, DGKG, GPR158, KCNC2
## PC_ 2
## Positive: CAMKK1, DGKQ, NT5DC3, CA7, ABCG4, HTR1A, C5orf28, OTOP2, HYKK, DPEP2
## CHST6, POTEI, SLC8A3, SLC38A11, ADRA2A, MPPED1, MTMR8, HTR7, CACNA1B, PPAPDC2
## C2orf69, GRIK1, IFI30, STK32B, RASL10B, SLC24A4, FAXDC2, ADCY3, ACSS2, ANKRD29
## Negative: RAN, HSP90AA1, H2AFZ, HNRNPAB, CCT5, NPM1, GNG5, DBI, HMGB2, ITM2B
## ATP6V1G1, SERPINH1, CIRBP, CD63, NDUFA6, MDK, JUN, MYL12B, SPARC, NPC2
## GLUL, ID3, EEF1A1, VIM, CLIC1, COX6B1, LDHA, DDAH2, ENO1, CNN3
## PC_ 3
## Positive: ADGRL2, AC011288.2, RP11-420N3.3, RP11-191L9.4, NRXN3, PLPPR1, RP11-123O10.4, ZNF385D, AC114765.1, NWD2
## RBFOX3, MIR137HG, MIR325HG, SGOL1-AS1, POU6F2, ANKRD18A, LY86-AS1, LINC01197, DGCR5, DPY19L1P1
## MIR4300HG, AQP4-AS1, HPSE2, LINC00632, NLGN4X, AC067956.1, PWRN1, LINC00599, CABP1, LINC01158
## Negative: KRTCAP2, APOE, C20orf24, PDIA6, PGLS, GNG11, S100A13, HIST1H2BI, ISCA2, GSTM5
## LAPTM4A, CST3, TMEM176B, KLF4, PDLIM2, CAP1, S100A16, APRT, CYR61, FAIM
## IFITM3, CDKN1A, KLF2, CLIC1, ARPC1B, IER2, S100A1, CMTM5, FXYD1, TCN2
## PC_ 4
## Positive: RESP18, CTXN2, ATP6V1G2, GNG13, DISP2, C15orf59, CCDC85A, GNG3, SYNGR3, RGS8
## VWA5B2, C1QL3, HPCA, TUBB3, CALB1, SNCB, HTR3A, ARHGDIG, L1CAM, NAP1L5
## PCDH20, HMP19, DBNDD2, NPAS4, FABP3, CALY, FAM43B, CKMT1B, LOC728392, LTK
## Negative: PTPN18, SLCO1A2, LINC00639, INPP5D, IFI44, LYN, DISC1, NEAT1, NRGN, CMYA5
## IFI44L, GALNT15, PARP14, AC012593.1, AQP4-AS1, MSR1, MT2A, ISG15, SHROOM4, CABP1
## UACA, KCNQ1OT1, PART1, CNDP1, FAM153B, DGCR5, SOX2-OT, LINC00844, ADGRG1, LINC00599
## PC_ 5
## Positive: MEST, IGFBP2, CNN3, FBXL7, NNAT, TUBB2B, GPC3, VIM, NKAIN4, ID1
## BMP7, CSRP2, NDN, DDAH2, GPX8, IGFBPL1, MARCKSL1, GSTM3, FBLN1, PARD3
## MFAP4, PTN, FABP7, COPS6, CTNNA2, ZBTB20, BEX1, CD81, ENO1, NPAS3
## Negative: C1QB, FCGR2A, MS4A6A, TYROBP, C1QC, AIF1, C1QA, CSF1R, CD86, MRC1
## MS4A7, CTSS, CCL24, FCER1G, CD53, CD14, FCGR1A, PLEK, C3AR1, LYZ
## FCGR2B, CX3CR1, CCL3L3, CCL2, CCR1, CD68, C5AR1, PF4, HPGDS, LY86## No graphs detected. Computing.## Computing nearest neighbor graph## Computing SNN## Using graph: RNA_snn## + Filtering results by `var1_search`: astro## + 6 entries matching `var1_search` identified.## + Adding original names to results## + Returning 50 pair-wise similarities.
cat("\nLabels similar to 'astro':\n")##
## Labels similar to 'astro':## [1] "fly.Aerts2020.Cortex_glia"
## [2] "fly.Aerts2020.Ensheathing_glia"
## [3] "fly.Aerts2020.Perineurial_glia"
## [4] "mouse.Zeisel2016.Astro2"
## [5] "mouse.Zeisel2016.Astro1"
## [6] "mouse.Zeisel2018.Subventricular.zone.radial.glia.like.cells"
## [7] "fly.Aerts2020.X76"
## [8] "fly.Aerts2020.Plasmatocytes"
## [9] "fly.Aerts2020.Subperineurial_glia"
## [10] "human.descartes_SampledData.Lens_fibre_cells"
## [11] "mouse.Zeisel2016.Oligo1"
## [12] "mouse.Zeisel2018.Olfactory.ensheathing.cells"
## [13] "human.descartes_SampledData.Inhibitory_neurons"
## [14] "human.descartes_SampledData.Amacrine_cells"
## [15] "human.descartes_SampledData.Horizontal_cells"
## [16] "mouse.Zeisel2018.Dentate.gyrus.radial.glia.like.cells"
## [17] "mouse.Zeisel2018.Ependymal.cells"
## [18] "human.descartes_SampledData.PDE11A_FAM19A2_positive_cells"
## [19] "mouse.Zeisel2018.Astrocytes"
## [20] "mouse.DRONC_mouse.ASC1"
## [21] "mouse.DRONC_mouse.ASC2"
## [22] "mouse.DRONC_mouse.ChP"
## [23] "mouse.Zeisel2016.Epend"
## [24] "mouse.DRONC_mouse.ODC"
## [25] "human.DRONC_human.NSC"
## [26] "mouse.Zeisel2016.Oligo2"
## [27] "mouse.Zeisel2016.Oligo6"
## [28] "mouse.Zeisel2018.Oligodendrocyte.precursor.cells"
## [29] "mouse.Zeisel2018.Oligodendrocytes"
## [30] "human.DRONC_human.ASC1"
## [31] "human.DRONC_human.ASC2"Strategy 3: Combined Approach
Combine TF-IDF with neighbor analysis for robust harmonization.
Identify Clusters by Enriched Terms
# Find clusters enriched for neuronal terms
neuronal_terms <- c("neuron", "neuronal", "glutamatergic", "gabaergic")
neuronal_clusters <- tfidf_df |>
dplyr::filter(word %in% neuronal_terms) |>
dplyr::group_by(cluster) |>
dplyr::summarise(
neuronal_score = sum(tf_idf),
top_term = word[which.max(tf_idf)]
) |>
dplyr::arrange(dplyr::desc(neuronal_score))## Warning: Returning more (or less) than 1 row per `summarise()` group was deprecated in
## dplyr 1.1.0.
## ℹ Please use `reframe()` instead.
## ℹ When switching from `summarise()` to `reframe()`, remember that `reframe()`
## always returns an ungrouped data frame and adjust accordingly.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.## `summarise()` has grouped output by 'cluster'. You can override using the
## `.groups` argument.
print(neuronal_clusters)## # A tibble: 3 × 3
## # Groups: cluster [15]
## cluster neuronal_score top_term
## <fct> <dbl> <chr>
## 1 6 0.0704 neuron
## 2 0 0.0428 neuron
## 3 2 0.0419 neuronValidate with Neighbor Search
Use neighbor search to validate that cells in these clusters are transcriptomically similar:
# Get cells from a neuronal cluster
if (nrow(neuronal_clusters) > 0) {
top_cluster <- neuronal_clusters$cluster[1]
is_top <- pseudo_seurat$cluster == top_cluster
cluster_cells <- pseudo_seurat$celltype[is_top]
cat("Labels in top neuronal cluster:\n")
print(table(cluster_cells))
}## Labels in top neuronal cluster:
## cluster_cells
## A
## 0
## AFP_ALB_positive_cells
## 0
## ASC1
## 0
## ASC2
## 0
## Acinar_cells
## 0
## Adrenocortical_cells
## 0
## Amacrine_cells
## 0
## Antigen_presenting_cells
## 0
## Ast
## 0
## AstA
## 0
## Astro1
## 0
## Astro2
## 0
## Astrocyte
## 0
## Astrocytes
## 0
## Bipolar_cells
## 0
## Blood_e7_0
## 0
## Blood_e8_0
## 0
## Blood_e8_5
## 0
## Blood_e9_0
## 0
## Blood_e10_0
## 0
## Blood_e11_0
## 0
## Blood_e12_0
## 0
## Blood_e12_5
## 0
## Blood_e13_0
## 0
## Blood_e13_5
## 0
## Blood_e14_0
## 0
## Blood_e14_5
## 0
## Blood_e15_0
## 0
## Blood_e15_5
## 0
## Blood_e16_0
## 0
## Blood_e16_5
## 0
## Blood_e16_25
## 0
## Blood_e17_0
## 0
## Blood_e17_5
## 0
## Blood_e18_0
## 0
## Bronchiolar_and_alveolar_epithelial_cells
## 0
## C2
## 0
## C3
## 0
## CA1Pyr1
## 0
## CA1Pyr2
## 0
## CA1PyrInt
## 0
## CA2Pyr2
## 0
## CCAP
## 0
## CCHa1
## 0
## CCL19_CCL21_positive_cells
## 0
## CLC_IL5RA_positive_cells
## 0
## CSH1_CSH2_positive_cells
## 0
## Capa
## 0
## Cardiomyocytes
## 0
## Cerebellum
## 1
## ChP
## 0
## Chiasm_glia
## 0
## Cholinergic
## 1
## Choroid
## 0
## Choroid_plexus_e9_0
## 0
## Choroid_plexus_e10_0
## 0
## Choroid_plexus_e11_0
## 0
## Choroid_plexus_e12_0
## 0
## Choroid_plexus_e12_5
## 0
## Choroid_plexus_e13_0
## 0
## Choroid_plexus_e13_5
## 0
## Choroid_plexus_e14_0
## 0
## Choroid_plexus_e14_5
## 0
## Choroid_plexus_e15_0
## 0
## Choroid_plexus_e15_5
## 0
## Choroid_plexus_e16_0
## 0
## Choroid_plexus_e16_5
## 0
## Choroid_plexus_e16_25
## 0
## Choroid_plexus_e17_0
## 0
## Choroid_plexus_e17_5
## 0
## Choroid_plexus_e18_0
## 0
## Chromaffin_cells
## 0
## Ciliated_epithelial_cells
## 0
## ClauPyr
## 0
## Clock
## 0
## Corneal_and_conjunctival_epithelial_cells
## 0
## Cortex_glia
## 0
## Crz
## 0
## DCN
## 0
## DN1
## 0
## Dentate
## 0
## Di
## 2
## Dm8
## 0
## Dm9
## 0
## Dopaminergic
## 0
## Ductal_cells
## 0
## ELF3_AGBL2_positive_cells
## 0
## END
## 0
## ENS_glia
## 0
## ENS_neurons
## 1
## Early_Neural_Tube_e7_0
## 0
## Early_Neural_Tube_e8_0
## 0
## Early_Neural_Tube_e8_5
## 0
## Ectoderm_e7_0
## 0
## Ectoderm_e8_0
## 0
## Ectoderm_e8_5
## 0
## Ectoderm_e9_0
## 0
## Ectoderm_e10_0
## 0
## Ectoderm_e11_0
## 0
## Ectoderm_e12_0
## 0
## Ectoderm_e12_5
## 0
## Ectoderm_e13_5
## 0
## Ectoderm_e14_5
## 0
## Ectoderm_e15_0
## 0
## Ectoderm_e15_5
## 0
## End
## 0
## Endocardial_cells
## 0
## Endoderm_e7_0
## 0
## Endoderm_e8_0
## 0
## Endoderm_e8_5
## 0
## Ensheathing_glia
## 0
## Enteric
## 1
## Epend
## 0
## Ependymal
## 0
## Epicardial_fat_cells
## 0
## Erythroblasts
## 0
## Ex1
## 0
## Ex2
## 0
## Ex3a
## 0
## Ex3b
## 0
## Ex3c
## 0
## Ex3d
## 0
## Ex3e
## 0
## Ex4
## 0
## Ex5a
## 0
## Ex5b
## 0
## Ex6a
## 0
## Ex6b
## 0
## Ex8
## 0
## Excitatory_neurons
## 0
## Extravillous_trophoblasts
## 0
## FMRFa
## 0
## Fibroblast_e9_0
## 0
## Fibroblast_e10_0
## 0
## Fibroblast_e11_0
## 0
## Fibroblast_e12_0
## 0
## Fibroblast_e12_5
## 0
## Fibroblast_e13_0
## 0
## Fibroblast_e13_5
## 0
## Fibroblast_e14_0
## 0
## Fibroblast_e14_5
## 0
## Fibroblast_e15_0
## 0
## Fibroblast_e15_5
## 0
## Fibroblast_e16_0
## 0
## Fibroblast_e16_5
## 0
## Fibroblast_e16_25
## 0
## Fibroblast_e17_0
## 0
## Fibroblast_e17_5
## 0
## Fibroblast_e18_0
## 0
## G
## 0
## GABA1
## 0
## GABA2
## 0
## GABAergic
## 0
## Ganglion_cells
## 0
## Glia_e9_0
## 0
## Glia_e10_0
## 0
## Glia_e11_0
## 0
## Glia_e12_0
## 0
## Glia_e12_5
## 0
## Glia_e13_0
## 0
## Glia_e13_5
## 0
## Glia_e14_0
## 0
## Glia_e14_5
## 0
## Glia_e15_0
## 0
## Glia_e15_5
## 0
## Glia_e16_0
## 0
## Glia_e16_5
## 0
## Glia_e16_25
## 0
## Glia_e17_0
## 0
## Glia_e17_5
## 0
## Glia_e18_0
## 0
## Glutamatergic
## 0
## Goblet_cells
## 0
## Gr43a
## 0
## Granule_neurons
## 0
## Hematopoietic_stem_cells
## 0
## Hepatoblasts
## 0
## Hindbrain
## 1
## Horizontal_cells
## 0
## Hsp
## 0
## Hug
## 0
## IGFBP1_DKK1_positive_cells
## 0
## IPC
## 0
## ITP
## 0
## Immune_e9_0
## 0
## Immune_e10_0
## 0
## Immune_e11_0
## 0
## Immune_e12_0
## 0
## Immune_e12_5
## 0
## Immune_e13_0
## 0
## Immune_e13_5
## 0
## Immune_e14_0
## 0
## Immune_e14_5
## 0
## Immune_e15_0
## 0
## Immune_e15_5
## 0
## Immune_e16_0
## 0
## Immune_e16_5
## 0
## Immune_e16_25
## 0
## Immune_e17_0
## 0
## Immune_e17_5
## 0
## Immune_e18_0
## 0
## In1a
## 0
## In1b
## 0
## In1c
## 0
## In2
## 0
## In3
## 0
## In4a
## 0
## In4b
## 0
## In6a
## 0
## In6b
## 0
## In7
## 0
## In8
## 0
## Inhibitory_interneurons
## 1
## Inhibitory_neurons
## 0
## Int1
## 0
## Int2
## 0
## Int3
## 0
## Int4
## 0
## Int5
## 0
## Int6
## 0
## Int7
## 0
## Int8
## 0
## Int9
## 0
## Int10
## 0
## Int11
## 0
## Int12
## 0
## Int13
## 0
## Int14
## 0
## Int15
## 0
## Int16
## 0
## Intestinal_epithelial_cells
## 0
## Islet_endocrine_cells
## 0
## L1
## 0
## L2
## 0
## L3
## 0
## L4
## 0
## LNv
## 0
## Lamina_monopolar
## 0
## Lawf1
## 0
## Lawf2
## 0
## Lens_fibre_cells
## 0
## Limbic_system_neurons
## 0
## Lymphatic_endothelial_cells
## 0
## Lymphoid_cells
## 0
## MBON
## 0
## MG
## 0
## MUC13_DMBT1_positive_cells
## 0
## Megakaryocytes
## 0
## Mesangial_cells
## 0
## Mesoderm_e7_0
## 0
## Mesoderm_e8_0
## 0
## Mesoderm_e8_5
## 0
## Mesothelial_cells
## 0
## Metanephric_cells
## 0
## Mgl1
## 0
## Mgl2
## 0
## Mi1
## 0
## Mic
## 0
## Microglia
## 0
## Mip
## 0
## Myeloid_cells
## 0
## NSC
## 0
## Neural_crest_e7_0
## 0
## Neural_crest_e8_0
## 0
## Neural_crest_e8_5
## 0
## Neural_crest_e9_0
## 0
## Neural_crest_e10_0
## 0
## Neural_crest_e11_0
## 0
## Neuroendocrine_cells
## 0
## Neuron_e8_0
## 0
## Neuron_e8_5
## 0
## Neuron_e9_0
## 0
## Neuron_e10_0
## 0
## Neuron_e11_0
## 0
## Neuron_e12_0
## 0
## Neuron_e12_5
## 1
## Neuron_e13_0
## 1
## Neuron_e13_5
## 1
## Neuron_e14_0
## 1
## Neuron_e14_5
## 0
## Neuron_e15_0
## 0
## Neuron_e15_5
## 0
## Neuron_e16_0
## 1
## Neuron_e16_5
## 0
## Neuron_e16_25
## 0
## Neuron_e17_0
## 0
## Neuron_e17_5
## 0
## Neuron_e18_0
## 0
## Node_e7_0
## 0
## Node_e8_0
## 0
## Non
## 0
## ODC
## 0
## ODC1
## 0
## ODC2
## 0
## OPC
## 0
## OPCs_e12_0
## 0
## OPCs_e12_5
## 0
## OPCs_e13_0
## 0
## OPCs_e13_5
## 0
## OPCs_e14_0
## 0
## OPCs_e14_5
## 0
## OPCs_e15_0
## 0
## OPCs_e15_5
## 0
## OPCs_e16_0
## 0
## OPCs_e16_5
## 0
## OPCs_e16_25
## 0
## OPCs_e17_0
## 0
## OPCs_e17_5
## 0
## OPCs_e18_0
## 0
## Octopaminergic
## 0
## Olfactory
## 0
## Olfactory_projection_neurons
## 0
## Oli
## 0
## Oligo1
## 0
## Oligo2
## 0
## Oligo3
## 0
## Oligo4
## 0
## Oligo5
## 0
## Oligo6
## 0
## Oligo_precursor
## 0
## Oligodendrocyte
## 0
## Oligodendrocytes
## 0
## PAEP_MECOM_positive_cells
## 0
## PAM
## 0
## PDE1C_ACSM3_positive_cells
## 0
## PDE11A_FAM19A2_positive_cells
## 0
## Parietal_and_chief_cells
## 0
## Peptidergic
## 1
## Per
## 0
## Peric
## 0
## Pericytes
## 0
## Perineurial_glia
## 0
## Peripheral
## 3
## Perivascular
## 0
## Photoreceptor_cells
## 0
## Photoreceptors
## 0
## Plasmatocytes
## 0
## Pluripotent_stem_cells_e7_0
## 0
## Pluripotent_stem_cells_e8_0
## 0
## Pluripotent_stem_cells_e8_5
## 0
## Pm1
## 0
## Pm3
## 0
## Pm4
## 0
## Poxn
## 0
## Proc
## 0
## Purkinje_neurons
## 0
## Pvm1
## 0
## Pvm2
## 0
## RPE
## 0
## RPE_differentiation
## 0
## Radial_glia_e9_0
## 0
## Radial_glia_e10_0
## 0
## Radial_glia_e11_0
## 0
## Radial_glia_e12_0
## 0
## Radial_glia_e12_5
## 0
## Radial_glia_e13_0
## 0
## Radial_glia_e13_5
## 0
## Radial_glia_e14_0
## 0
## Radial_glia_e14_5
## 0
## Radial_glia_e15_0
## 0
## Radial_glia_e15_5
## 0
## Radial_glia_e16_0
## 0
## Radial_glia_e16_5
## 0
## Radial_glia_e16_25
## 0
## Radial_glia_e17_0
## 0
## Radial_glia_e17_5
## 0
## Radial_glia_e18_0
## 0
## Retinal_pigment_cells
## 0
## Retinal_progenitors_and_Muller_glia
## 0
## S1PyrDL
## 0
## S1PyrL4
## 0
## S1PyrL5
## 0
## S1PyrL5a
## 0
## S1PyrL6
## 0
## S1PyrL6b
## 0
## S1PyrL23
## 0
## SATB2_LRRC7_positive_cells
## 0
## SKOR2_NPSR1_positive_cells
## 0
## SLC24A4_PEX5L_positive_cells
## 0
## SLC26A4_PAEP_positive_cells
## 0
## SMC
## 0
## STC2_TLX1_positive_cells
## 0
## Satellite
## 0
## Satellite_cells
## 0
## Schwann_cells
## 0
## Schwann_e12_0
## 0
## Schwann_e13_0
## 0
## Schwann_e13_5
## 0
## Schwann_e14_0
## 0
## Schwann_e14_5
## 0
## Schwann_e15_0
## 0
## Schwann_e15_5
## 0
## Schwann_e16_0
## 0
## Schwann_e16_5
## 0
## Schwann_e16_25
## 0
## Schwann_e17_0
## 0
## Schwann_e17_5
## 0
## Schwann_e18_0
## 0
## Serotonergic
## 0
## Skeletal_muscle_cells
## 0
## Smooth_muscle_cells
## 0
## Spinal
## 2
## Squamous_epithelial_cells
## 0
## Stellate_cells
## 0
## Stromal_cells
## 0
## SubPyr
## 0
## Subperineurial_glia
## 0
## Subventricular
## 0
## Sympathetic
## 2
## Sympathoblasts
## 0
## Syncytiotrophoblasts_and_villous_cytotrophoblasts
## 0
## T1
## 0
## T2
## 0
## T3
## 0
## T4
## 0
## Telencephalon
## 0
## Thymic_epithelial_cells
## 0
## Thymocytes
## 0
## Tm1
## 0
## Tm5ab
## 0
## Tm5c
## 0
## Tm9
## 0
## TmY14
## 0
## Trophectoderm_e7_0
## 0
## Trophectoderm_e8_0
## 0
## Trophectoderm_e8_5
## 0
## Trophoblast_giant_cells
## 0
## Tyraminergic
## 0
## Unipolar_brush_cells
## 0
## Ureteric_bud_cells
## 0
## Vascular
## 0
## Vascular_e7_0
## 0
## Vascular_e8_0
## 0
## Vascular_e8_5
## 0
## Vascular_e9_0
## 0
## Vascular_e10_0
## 0
## Vascular_e11_0
## 0
## Vascular_e12_0
## 0
## Vascular_e12_5
## 0
## Vascular_e13_0
## 0
## Vascular_e13_5
## 0
## Vascular_e14_0
## 0
## Vascular_e14_5
## 0
## Vascular_e15_0
## 0
## Vascular_e15_5
## 0
## Vascular_e16_0
## 0
## Vascular_e16_5
## 0
## Vascular_e16_25
## 0
## Vascular_e17_0
## 0
## Vascular_e17_5
## 0
## Vascular_e18_0
## 0
## Vascular_endothelial_cells
## 0
## Vend1
## 0
## Vend2
## 0
## Visceral_neurons
## 1
## Vsmc
## 0
## X
## 0
## X0
## 0
## X1
## 0
## X2
## 0
## X3
## 0
## X4
## 0
## X5
## 0
## X6
## 0
## X7
## 0
## X9
## 0
## X13
## 0
## X15
## 0
## X16
## 0
## X18
## 0
## X19
## 0
## X21
## 0
## X25
## 0
## X29
## 0
## X30
## 0
## X32
## 0
## X34
## 0
## X40
## 0
## X44
## 0
## X47
## 0
## X49
## 0
## X50
## 0
## X51
## 0
## X54
## 0
## X56
## 0
## X59
## 0
## X62
## 0
## X67
## 0
## X68
## 0
## X69
## 0
## X71
## 0
## X75
## 0
## X76
## 0
## X78
## 0
## X79
## 0
## X81
## 0
## X83
## 0
## X84
## 0
## adPN
## 0
## adaxial_cell
## 0
## blood_precursors
## 0
## cartilage
## 0
## cerebellum
## 0
## comitted_progenitors
## 0
## comittted_progenitors
## 0
## committed_progenitors
## 0
## cornea
## 0
## cranial_ganglion
## 2
## cycling_cells
## 0
## cycling_progenitors
## 0
## dermal_bone
## 0
## dienc
## 0
## diencephalon
## 1
## dorsal_Fan
## 0
## dorsal_diencephalon
## 0
## dorsal_diencephalon_II
## 0
## dorsal_dienchephalon
## 0
## dorsal_habenula
## 2
## dorsal_telencephalon
## 2
## epidermal_ionocyte
## 0
## epidermis
## 0
## epithelium
## 0
## erythrocytes
## 0
## exCA1
## 0
## exCA3
## 0
## exDG
## 0
## exPFC1
## 0
## exPFC2
## 0
## exPFC3
## 0
## exPFC4
## 0
## exPFC5
## 0
## exPFC6
## 0
## exPFC7
## 0
## exPFC8
## 0
## exPFC9
## 0
## eye
## 0
## floor_plate
## 0
## glia
## 0
## glia_progenitors
## 0
## glial_progenitors
## 0
## granule_cells
## 1
## heart_primordium
## 0
## hinbrain_rhombomere_5
## 0
## hinbrain_rhombomere_6
## 0
## hinbrain_rhombomere_7
## 0
## hindbrain
## 0
## hindbrain_rhombomere_1
## 0
## hindbrain_rhombomere_2
## 0
## hindbrain_rhombomere_3
## 0
## hindbrain_rhombomere_4
## 0
## hindbrain_rhombomere_5
## 0
## hindbrain_rhombomere_6
## 0
## hindbrain_rhombomere_7
## 0
## hypothalamus
## 1
## lPN
## 0
## lens
## 0
## lens_differentiation
## 0
## lens_epithelium
## 0
## macrophages
## 0
## mesoderm
## 0
## microglia
## 0
## mid
## 1
## midbrain
## 1
## midbrain_neural_rod
## 0
## muscle
## 0
## neural_crest
## 0
## neural_crest_derived
## 0
## neuromast
## 0
## neurons
## 12
## neutrophils
## 0
## notochord
## 0
## olfactory_bulb
## 0
## oligodendrocytes
## 0
## optic_stalk
## 0
## optic_tectum
## 1
## optic_vesicle
## 0
## otic
## 0
## otic_vesicle
## 0
## pectoral_fin
## 0
## pectoral_fin_field
## 0
## periderm
## 0
## pharangeal_arch
## 0
## pharyneal_arch
## 0
## pharyngeal_arch
## 0
## pharyngeal_endoderm
## 0
## pharyngeal_erythrocyte_lineage
## 0
## pigment_cell
## 0
## placode
## 0
## possibly_otic
## 0
## prechordal_plate
## 0
## progenitors
## 1
## pronephros
## 0
## purkinje_neurons
## 2
## radial_glia
## 0
## retina
## 2
## retina_neuroblasts
## 0
## retina_neurogenesis
## 0
## retina_neurons
## 1
## rostral_blood_island
## 0
## sensory_neurons
## 0
## somite
## 0
## telencephalon
## 2
## telencephalon_I
## 0
## telencephalon_II
## 0
## ventral_diencephalon
## 0
## ventral_forebrain
## 0
## ventral_habenula
## 0
## ventral_midbrain
## 0Visualizing Cross-dataset Agreement
Color by Dataset
Visualize how cells from different datasets intermix:
res_batch <- plot_tfidf(
obj = pseudo_seurat,
label_var = "celltype",
cluster_var = "cluster",
color_var = "batch",
show_plot = TRUE
)## Extracting obsm from Seurat: umap## + Dropping 2 conflicting obs variables: UMAP.1, UMAP.2## Setting cell metadata (obs) in obj.## Warning in ggplot2::geom_point(ggplot2::aes_string(color = color_var, size =
## size_var, : Ignoring unknown aesthetics: label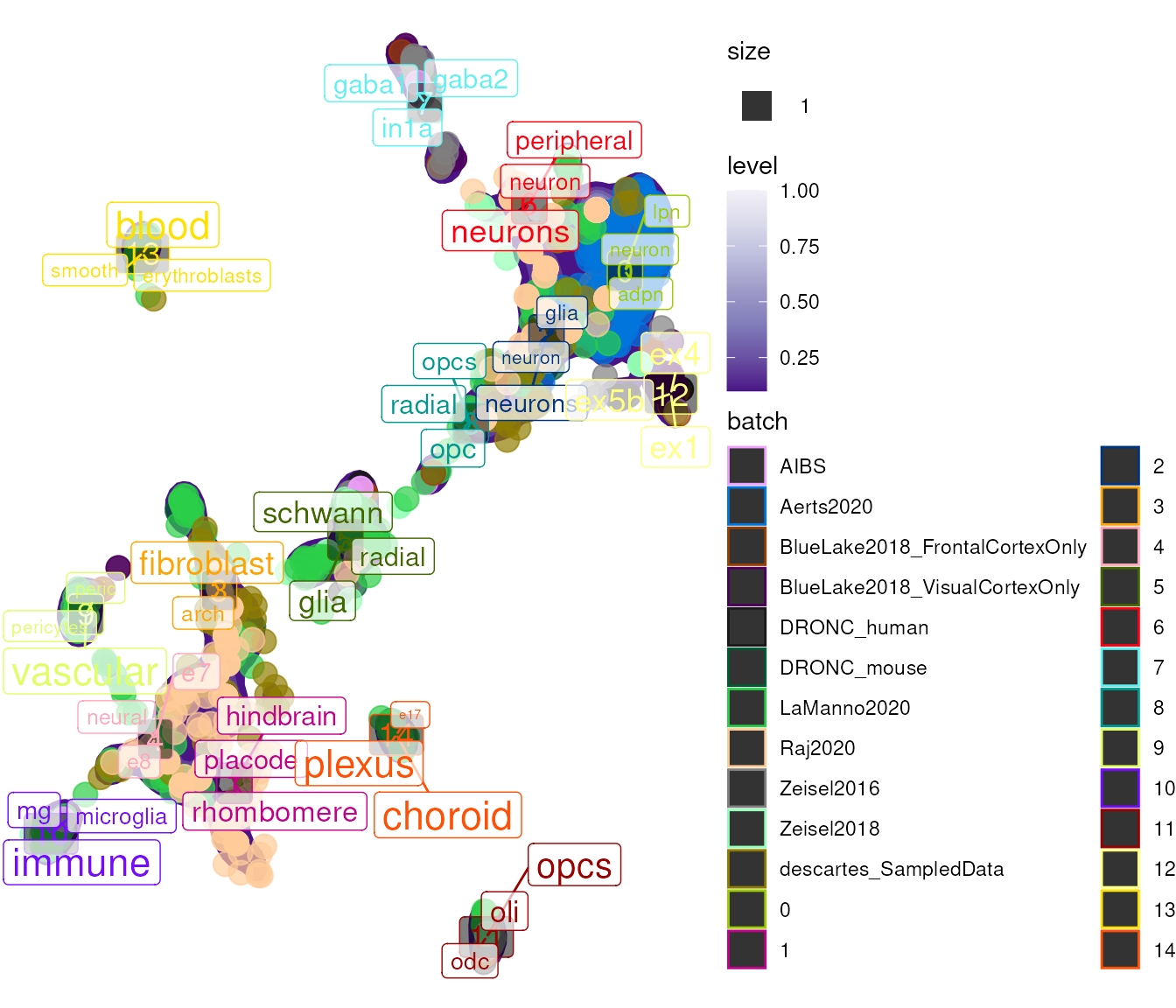
Clusters with cells from multiple datasets that share enriched terms indicate successful cross-dataset harmonization.
Color by Species
For cross-species comparisons:
res_species <- plot_tfidf(
obj = pseudo_seurat,
label_var = "celltype",
cluster_var = "cluster",
color_var = "species",
show_plot = TRUE
)## Extracting obsm from Seurat: umap## + Dropping 2 conflicting obs variables: UMAP.1, UMAP.2## Setting cell metadata (obs) in obj.## Warning in ggplot2::geom_point(ggplot2::aes_string(color = color_var, size =
## size_var, : Ignoring unknown aesthetics: label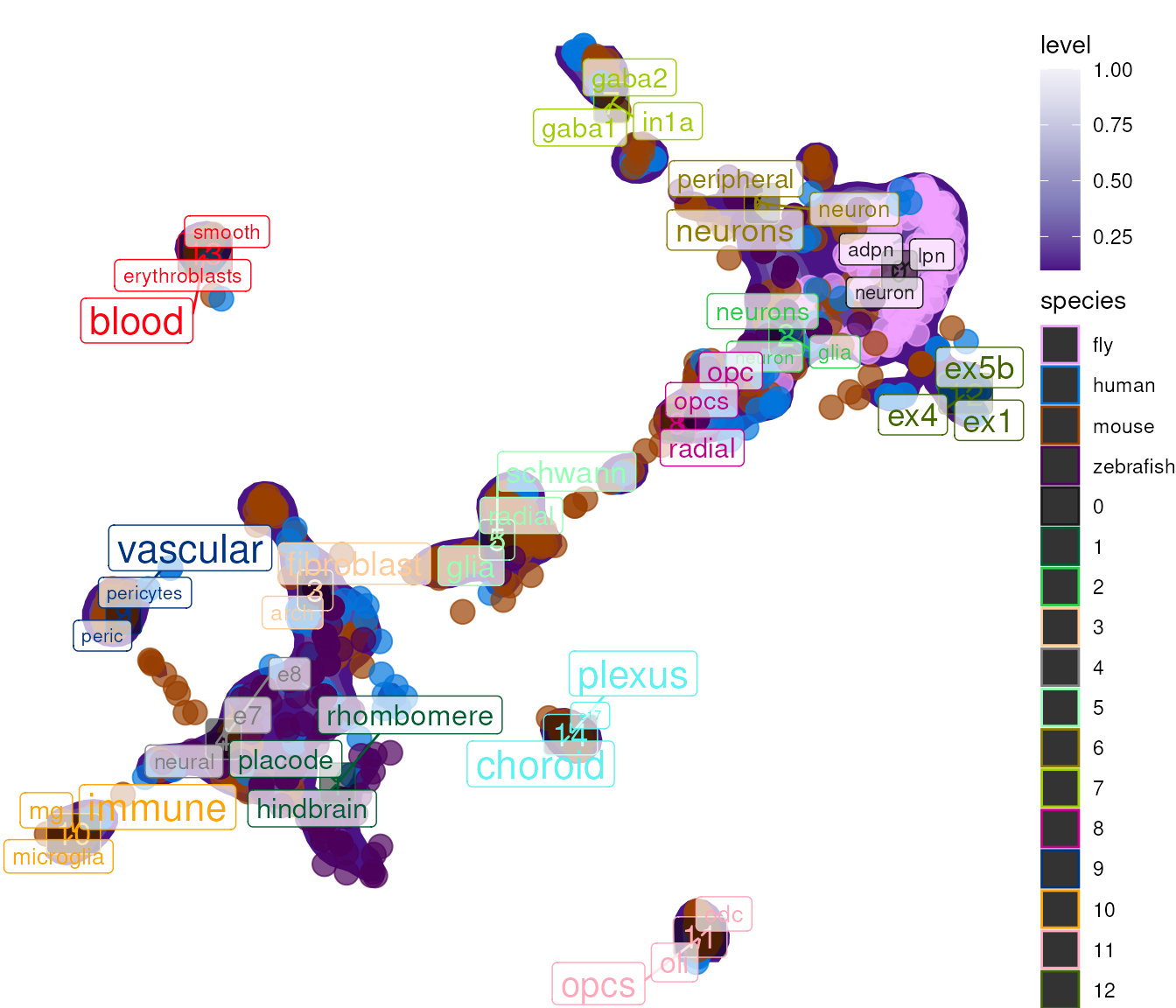
Word Cloud Summary
Generate a word cloud to summarize the most distinctive terms across all clusters:
wc <- wordcloud_tfidf(
obj = pseudo_seurat,
label_var = "celltype",
cluster_var = "cluster",
terms_per_cluster = 5
)## Loading required namespace: ggwordcloud## Extracting obsm from Seurat: umap## + Dropping 2 conflicting obs variables: UMAP.1, UMAP.2## Setting cell metadata (obs) in obj.## Warning in ggplot2::geom_point(ggplot2::aes_string(color = color_var, size =
## size_var, : Ignoring unknown aesthetics: label
Practical Recommendations
Start with TF-IDF: Identify enriched terms to understand the semantic content of each cluster.
Use neighbor search for validation: Confirm that cells with similar labels are transcriptomically similar.
Examine cross-dataset mixing: Well-harmonized cell types should have cells from multiple datasets in the same cluster.
Iterate: Refine cluster assignments and re-run TF-IDF to improve harmonization.
Document mappings: Create a mapping table of original labels to harmonized labels for reproducibility.
Session Info
utils::sessionInfo()## R Under development (unstable) (2026-01-22 r89323)
## Platform: x86_64-pc-linux-gnu
## Running under: Ubuntu 24.04.3 LTS
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
## LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
##
## locale:
## [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
## [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
## [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
##
## time zone: UTC
## tzcode source: system (glibc)
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] future_1.69.0 scNLP_0.99.0 BiocStyle_2.39.0
##
## loaded via a namespace (and not attached):
## [1] RColorBrewer_1.1-3 jsonlite_2.0.0 magrittr_2.0.4
## [4] spatstat.utils_3.2-1 farver_2.1.2 rmarkdown_2.30
## [7] fs_1.6.6 ragg_1.5.0 vctrs_0.7.1
## [10] ROCR_1.0-12 spatstat.explore_3.7-0 htmltools_0.5.9
## [13] janeaustenr_1.0.0 sass_0.4.10 sctransform_0.4.3
## [16] parallelly_1.46.1 KernSmooth_2.23-26 bslib_0.9.0
## [19] htmlwidgets_1.6.4 tokenizers_0.3.0 desc_1.4.3
## [22] ica_1.0-3 plyr_1.8.9 plotly_4.12.0
## [25] zoo_1.8-15 cachem_1.1.0 commonmark_2.0.0
## [28] igraph_2.2.1 mime_0.13 lifecycle_1.0.5
## [31] pkgconfig_2.0.3 Matrix_1.7-4 R6_2.6.1
## [34] fastmap_1.2.0 fitdistrplus_1.2-6 shiny_1.12.1
## [37] digest_0.6.39 tidytext_0.4.3 colorspace_2.1-2
## [40] patchwork_1.3.2 Seurat_5.4.0 tensor_1.5.1
## [43] RSpectra_0.16-2 irlba_2.3.5.1 SnowballC_0.7.1
## [46] textshaping_1.0.4 labeling_0.4.3 progressr_0.18.0
## [49] spatstat.sparse_3.1-0 httr_1.4.7 polyclip_1.10-7
## [52] abind_1.4-8 compiler_4.6.0 withr_3.0.2
## [55] S7_0.2.1 fastDummies_1.7.5 maps_3.4.3
## [58] MASS_7.3-65 tools_4.6.0 lmtest_0.9-40
## [61] otel_0.2.0 httpuv_1.6.16 future.apply_1.20.1
## [64] goftest_1.2-3 glue_1.8.0 nlme_3.1-168
## [67] promises_1.5.0 gridtext_0.1.5 grid_4.6.0
## [70] Rtsne_0.17 cluster_2.1.8.1 reshape2_1.4.5
## [73] generics_0.1.4 isoband_0.3.0 gtable_0.3.6
## [76] spatstat.data_3.1-9 tidyr_1.3.2 data.table_1.18.0
## [79] xml2_1.5.2 utf8_1.2.6 sp_2.2-0
## [82] spatstat.geom_3.7-0 RcppAnnoy_0.0.23 markdown_2.0
## [85] ggrepel_0.9.6 RANN_2.6.2 pillar_1.11.1
## [88] stringr_1.6.0 pals_1.10 spam_2.11-3
## [91] RcppHNSW_0.6.0 later_1.4.5 splines_4.6.0
## [94] dplyr_1.1.4 lattice_0.22-7 survival_3.8-6
## [97] deldir_2.0-4 tidyselect_1.2.1 miniUI_0.1.2
## [100] pbapply_1.7-4 knitr_1.51 gridExtra_2.3
## [103] litedown_0.9 bookdown_0.46 scattermore_1.2
## [106] xfun_0.56 matrixStats_1.5.0 stringi_1.8.7
## [109] lazyeval_0.2.2 yaml_2.3.12 evaluate_1.0.5
## [112] codetools_0.2-20 ggwordcloud_0.6.2 tibble_3.3.1
## [115] BiocManager_1.30.27 cli_3.6.5 uwot_0.2.4
## [118] xtable_1.8-4 reticulate_1.44.1 systemfonts_1.3.1
## [121] jquerylib_0.1.4 dichromat_2.0-0.1 Rcpp_1.1.1
## [124] globals_0.18.0 spatstat.random_3.4-4 mapproj_1.2.12
## [127] png_0.1-8 spatstat.univar_3.1-6 parallel_4.6.0
## [130] pkgdown_2.2.0 ggplot2_4.0.1 dotCall64_1.2
## [133] listenv_0.10.0 viridisLite_0.4.2 scales_1.4.0
## [136] ggridges_0.5.7 SeuratObject_5.3.0 purrr_1.2.1
## [139] rlang_1.1.7 cowplot_1.2.0